On-line storage
- DropBox, Google Drive, etc. alternative
- Encrypted
- Trustworthy third-parties
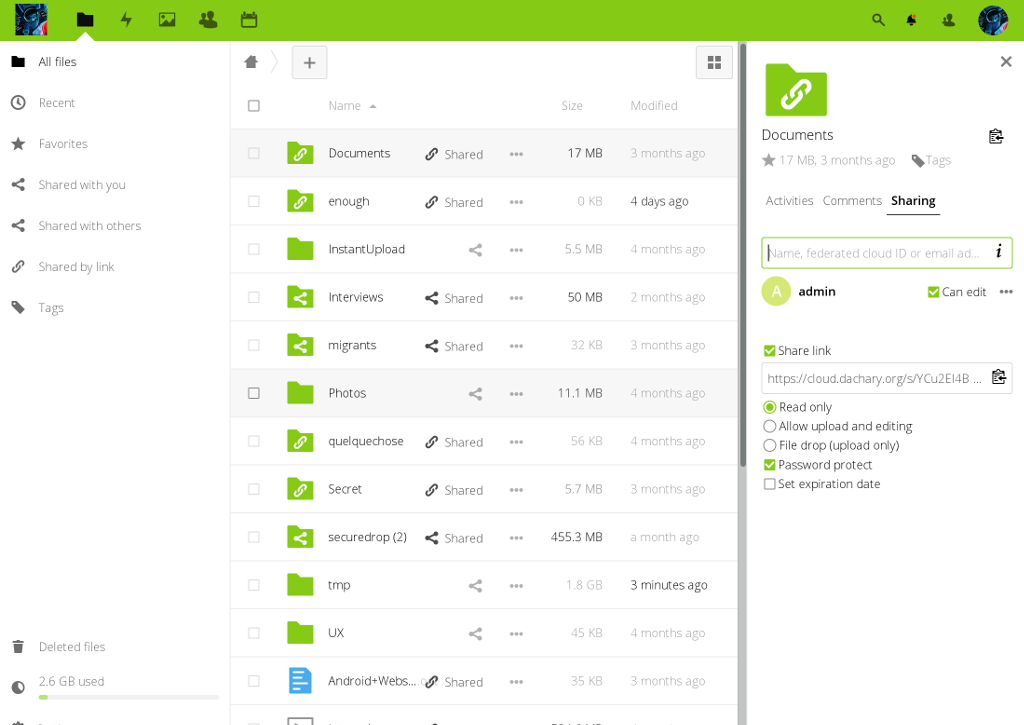
Self-Hosted & sustainable
- In the cloud or self-hosted
- On-premises
- Light weight maintenance

Copyright (c) 2022, Enough Community.
Template by Bootstrapious. Ported to Hugo by DevCows


